Chapter 6 Diagnostic microbiology and laboratory methods
Diagnostic microbiology
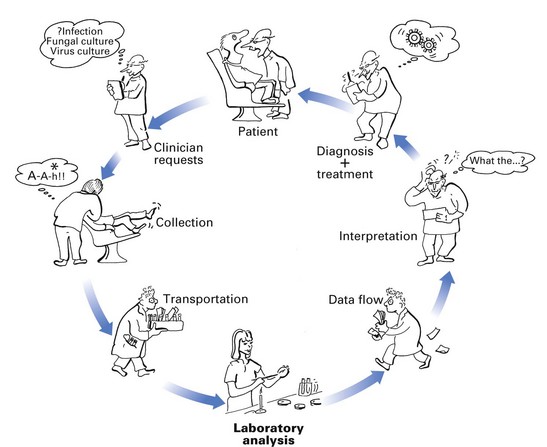
The diagnosis of an infectious disease entails a number of decisions and actions by many people. The diagnostic cycle begins when the clinician takes a microbiological sample and ends when the clinician receives the laboratory report and uses the information to manage the condition (Fig. 6.1). The steps in the diagnostic cycle are:
Clinical request
Laboratory analysis
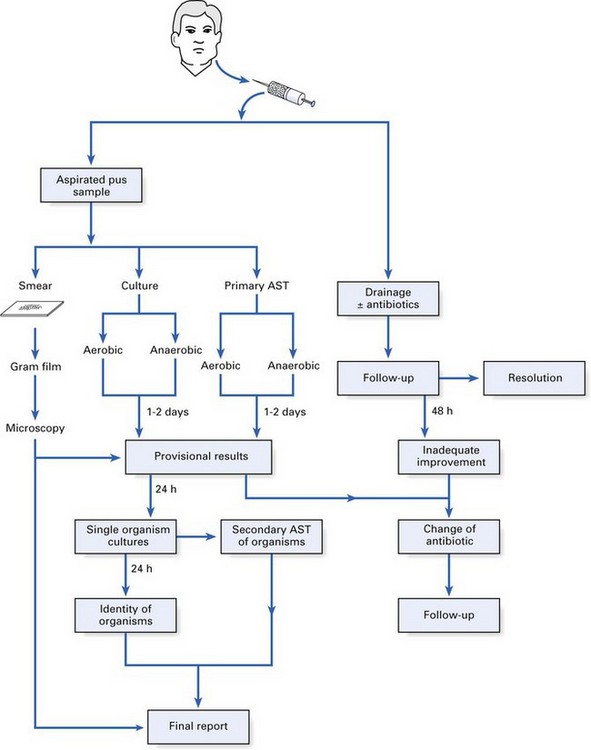
A wide array of specimens are received and analyzed by a number of methods in diagnostic microbiology laboratories. The analytical process of a pus specimen from a dental abscess is given below, as an illustration (Fig. 6.2):
Finally, it should be noted that the microbiologist can issue a provisional report after 2 days but the final report may take longer (Fig. 6.2).
Laboratory methods
Microscopic methods
Light microscopy
Light microscopy and stains
In light microscopy, bacterial stains are used:
The most commonly used stain in diagnostic microbiology is the Gram stain.
Gram stain technique
Detection of microbes by probing for their genes
Polymerase chain reaction
Very small bacterial numbers (10–100) in patient specimens can be detected using the standard polymerase chain reaction (PCR) techniques (Chapter 3), while more sophisticated techniques can detect one human immunodeficiency virus (HIV) proviral DNA sequence in 106 cells. The main advantage of this method is its rapidity (a few hours compared with many days for conventional cultural techniques). However, PCR reactions may yield non-specific data and hence judicial selection of primers and careful conduct of the assays (to prevent contaminants giving rise to false-positive results) are important. For these reasons, PCR techniques are not common in the diagnostic laboratory, but with new developments such as microarray technology and nested PCR, it is only a matter of time before this technique becomes more popular.
Cultural methods
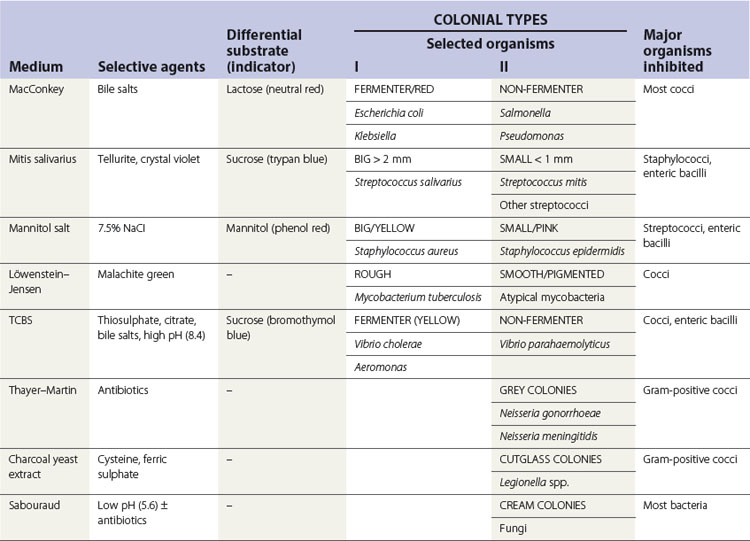
Bacteria grow well on artificial media, unlike viruses that require live cells for growth. Blood agar is the most widely used bacterial culture medium. It is an example of a non-selective medium as many organisms can grow on it. However, when chemicals are incorporated into media to prevent the growth of certain bacterial species and to promote the growth of others, selective media can be developed (e.g. the addition of bile salts helps the isolation of enterobacteria from a stool sample by suppressing the growth of most gut commensals). Some examples of selective media and their use are given in Table 6.1.
Bacteriological media
The main constituents of bacteriological media are:
Preparation of solid media and inoculation procedure
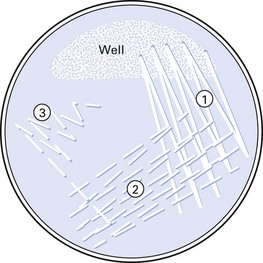
The objective of inoculating the specimen or a culture of bacteria on to a solid medium is to obtain discrete colonies of organisms after appropriate incubation. Hence, a standard technique (Fig. 6.3) should be used. Solid media are more useful than liquid media as they facilitate:
Liquid media
Liquid media are used in microbiology to:
Some examples of solid and liquid media are given in Table 6.2.
Table 6.2 Constituents and uses of some commonly used solid and liquid media
| Medium | Major ingredients | Use |
|---|---|---|
| Solid media | ||
| Nutrient agar | Nutrient broth, agar | General purpose |
| Blood agar | Nutrient agar, 5–10% horse or sheep blood | Very popular, general use |
| Chocolate agar | Heated blood agar | Isolation of Haemophilus and Neisseria spp. |
| CLED agar | Peptone, l-cystine, lactose, etc. | Culture of coliforms |
| Antibiotic sensitivity | Peptone and a semisynthetic medium | Antibiotic sensitivity tests |
| Liquid media | ||
| Peptone | Peptone, sodium chloride, water | General use; base for sugar fermentation tests |
| Nutrient broth | Peptone water, meat extract | General culture |
| Robertson’s meat medium | Nutrient broth, minced meat | Mainly to culture anaerobes |
| Selenite F broth | Peptone, water, sodium selenite | Enrichment medium for Salmonella and Shigella spp. |
CLED, cystine–lactose–electrolyte-deficient.
Media for blood culture
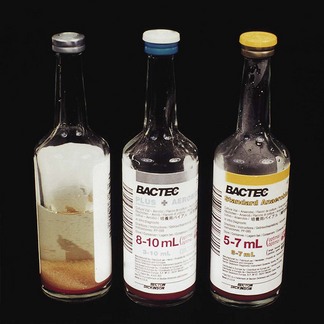
When the infectious agent is circulating in blood (e.g. in septicaemia, endocarditis, pneumonia), the latter has to be aseptically withdrawn by venepuncture and cultured. Blood culture has to be performed on special liquid media, under both aerobic and anaerobic conditions. The blood is aseptically transferred to a rich growth medium (e.g. brain–heart infusion broth) containing anticoagulants (Fig. 6.4). Cultures are checked for turbidity and gas production daily, up to a week (in many laboratories, this process is now automated and machines are used to detect bacterial growth). Positive cultures are sampled, and the organisms are isolated and identified.
Atmospheric requirements and incubation
Once inoculated, the agar plates may be incubated:
Stay updated, free dental videos. Join our Telegram channel

VIDEdental - Online dental courses