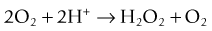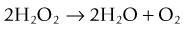Chapter 3 Bacterial physiology and genetics
Bacterial physiology
Growth
Nutritional requirements
Carbon
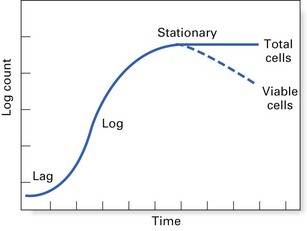
Bacterial growth cycle
The growth cycle of a bacterium has four main phases (Fig. 3.1):
Growth regulation
Extracellular factors that modify bacterial growth are:
 mesophiles, which grow well between 25 and 40°C, comprising most medically important bacteria (that grow best at body temperature)
mesophiles, which grow well between 25 and 40°C, comprising most medically important bacteria (that grow best at body temperature)Aerobic and anaerobic growth
and the second is catalase, which converts hydrogen peroxide to water and oxygen:
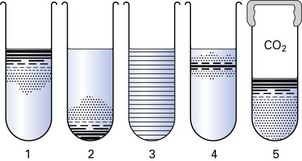
Bacteria can therefore be classified according to their ability to live in an oxygen-replete or an oxygen-free environment (Fig. 3.2, Table 3.1). This has important practical implications, as clinical specimens must be incubated in the laboratory under appropriate gaseous conditions for the pathogenic bacteria to grow. Thus, bacteria can be classified as follows:
| Degree of oxygenation | Term | Example |
|---|---|---|
| Oxygen essential for growth | Obligate aerobe | Pseudomonas aeruginosa |
| Grows well under low oxygen concentration (5%) | Microaerophile | Campylobacter fetus |
| Grows in the presence or absence of oxygen | Facultative anaerobea | Streptococcus milleri |
| Only grows in the absence of oxygen | Obligate anaerobe | Porphyromonas gingivalis |
a Facultative anaerobes may be subgrouped as capnophiles or capnophilic organisms if they grow well in the presence of 8–10% carbon dioxide (e.g. Legionella pneumophila).
Bacterial genetics
The bacterial chromosome
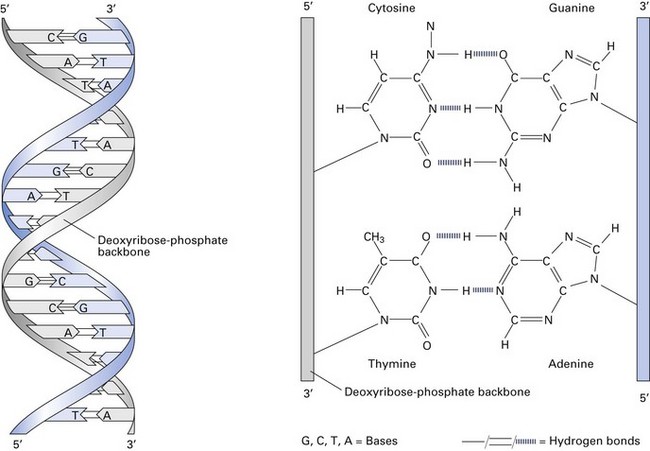
The bacterial chromosome contains the genetic information that defines all the characteristics of the organism. It is a single, continuous strand of DNA (Fig. 3.3) with a closed, circular structure attached to the cell membrane of the organism. The ‘average’ bacterial chromosome has a molecular weight of 2 × 109.
Replication
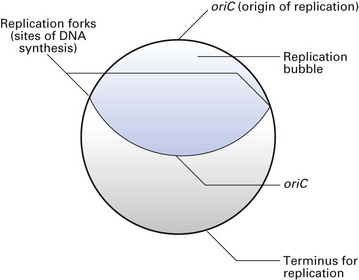
Chromosome replication is an accurate process that ensures that the progeny cells receive identical copies from the mother cell. The replication process is initiated at a specific site on the chromosome (oriC site) where the two DNA strands are locally denatured. A complex of proteins binds to this site, opens up the helix and initiates replication. Each strand then serves as a template for a complete round of DNA synthesis, which occurs in both directions (bidirectional) and on both strands, creating a replication bubble (Fig. 3.4). The two sites at which the replication occurs are called the replication forks. As replication proceeds, the replication forks move around the molecule in opposite directions opening up the DNA strands, synthesizing two new complementary strands until the two replication forks meet at a termination site. Of the four DNA strands now available, each daughter cell receives a parental strand and a newly synthesized strand. This process is called semiconservative replication. Such chromosomal replication is synchronous with cell division, so that each cell receives a full complement of DNA from the mother cell.
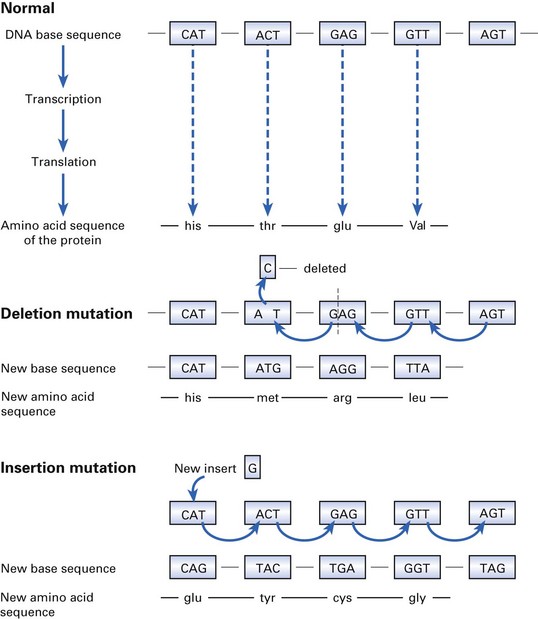
Genetic variation in bacteria
Genetic variation can occur as a result of mutation or gene transfer.
Stay updated, free dental videos. Join our Telegram channel

VIDEdental - Online dental courses