Introduction
The aim of this study was to evaluate biofilm retention around orthodontic brackets related to the method of ligation by using optical coherence tomography (OCT) and microbiologic sampling.
Methods
Seventy-five plastic central incisors for dentures were divided into 3 groups and used with metal brackets with a 0.022-in slot with elastomeric ligature (n = 25), metal brackets with a 0.022-in slot with steel wire ligature (n = 25), and self-ligating brackets with a 0.022-in slot (n = 25). The samples were submersed in a suspension of Streptococcus mutans , genetically engineered to express green fluorescent protein, at 37°C for 72 hours to allow biofilm formation. The samples were then submitted to microbiologic analysis and OCT imaging.
Results
The microbiologic analysis and the OCT showed significant differences in biofilm formation depending on the ligating method. Brackets ligated with elastomeric rings held more S mutans biofilm, and steel wire ligation had less biofilm retention compared with the other brackets.
Conclusions
This study provided validation that OCT can be used as a potential qualitative marker of total plaque bacteria that can be rapidly and reliably visualized around orthodontic brackets.
Orthodontic treatment with fixed appliances leads to increased biofilm accumulation and elevated levels of cariogenic and periodontal bacteria, mainly because orthodontic brackets make good oral hygiene difficult, resulting in plaque accumulation and significantly increased risks for enamel demineralization or periodontal disease.
Fixed orthodontic appliances create new retention areas that are suitable for bacterial colonization and lead to increases in the absolute numbers and the percentages of Streptococcus mutans and lactobacilli. Many studies have evaluated the effect of fixed orthodontic appliances on microbial flora and periodontal status ; however, only a few studies evaluated the method of ligation as an additional factor.
Conventionally, to connect the wire to the bracket, 3 methods are used: ligation with steel wires, elastomeric ligatures, and self-ligating brackets. All of these methods have advantages and disadvantages, but, concerning biofilm retention, the literature suggests that elastomeric ligatures increase the retention of dental biofilm compared with the 2 other methods.
In this study, we used a noninvasive method to visualize the biofilm formed around the brackets related to the ligating method: optical coherence tomography (OCT). This technique provides a 3-dimensional (3D) way to visualize the biofilm and allows a quantitative analysis of the retention area. For in-vitro studies, it can provide a reliable method for measuring biofilm area and thickness without destruction or alteration of the samples. Typically, the methods for plaque identification in vitro use microbiologic sampling or scanning electron microscopy that are invasive and destructive. In vivo, the procedures include biologic dyes and digital images to visualize and quantify the biofilm; however, this is normally uncomfortable for patients, since the dyes can stain oral tissues or orthodontic appliances. In addition, the dye has difficulty in staining thin biofilms. OCT does not need dyes to identify the biofilm and can provide image enhancement in a clinical environment that is impossible to obtain without a microscope.
OCT has been developed for cross-sectional imaging in biologic systems. It uses low-coherence interferometry and explores the optical properties of scattering samples to produce images from internal tissue microstructures in a way that is analogous to ultrasound pulse-echo imaging. This technique is a noninvasive, harmless diagnostic method and can depict the microstructure of biologic samples. The use of OCT to reveal biofilms on brackets is innovative. Image software can process the data obtained by OCT to quantify the biofilm retention area and identify in a 3D analysis the site of retention.
The aims of this study were to evaluate biofilm retention around orthodontic brackets as function of the method of ligation by using OCT and to compare the results with conventional microbiologic sampling.
Material and methods
Seventy-five plastic central incisors for dentures (Biotone, Dentsply, Rio de Janeiro, Brazil) were used. The samples were divided into 3 groups: conventional metal brackets with a 0.022-in slot with elastomeric ligature (n = 25), the same type of brackets with steel wires ligature (n = 25), and 25 self-ligating brackets with a 0.022-in slot for the straight-arch technique (respectively, Ovation and in-Ovation brackets; Dentsply/GAC, Bohemia, NY). All brackets used were maxillary central incisor brackets, bonded with composite bonding material (TPH, Dentsply). The samples were disinfected before the experiment by immersion in 1 mL of 70% ethanol for 30 minutes.
Genetically modified bacteria, S mutans (ATCC 25175), to express the green fluorescent protein were aerobically grown in brain-heart infusion broth at 37°C with shaking (150 rpm) for 48 hours, until they formed a stationary growth phase suspension. Genetically modified fluorescent bacteria were used to easily confirm, by image analysis, the formation of biofilms and the identification of the main site of biofilm retention on the brackets.
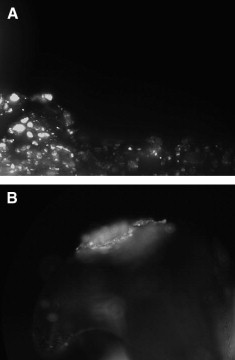
Bacterial cells were suspended in broth nutrient medium of brain-heart infusion to achieve a cell concentration of 10 9 colony-forming units per milliliter (confirmed by photospectroscopy at λ = 540 nm). One-milliliter aliquots were added to a petri dish with approximately 20 mL of brain-heart infusion broth with the samples inside. The petri dish was then sealed, kept upright, and incubated aerobically for 72 hours at 37°C with shaking (100 rpm) to allow biofilm formation. The brain-heart infusion broth was changed each 24 hours, and the presence of a microbial biofilm rather than planktonic bacteria was confirmed by the failure of irrigation with saline solution to significantly diminish the fluorescence signal analyzed at a wavelength of 520 nm by fluorescence microscopy ( Fig 1 ).
The configuration used in the OCT system was a fiber Michelson interferometer (OCP930SR; Thorlabs, Newton, NJ) that uses a superluminescent diode at 930 nm with 2 mW of optical power in the sample arm. The system can produce up to 8 frames per second with 2000 × 512 pixels with axial and lateral resolutions in air of 6.0 μm. The system has a hand probe implemented with optical fiber, making data collection easier.
For tridimensional image reconstruction, a precise displacement of the sample in the y-axis (orthogonal to the image acquisition by the OCT system) during the image acquisition was done by a computer-controlled linear translation stage (T25-XYZ, Thorlabs) with a minimum step of 0.05 μm. The 3D images were reconstructed by using VGStudioMax (version 1.2; Volume Graphics, Heidelberg, Germany) and analyzed with the software Image J (National Institutes of Health, Bethesda, Md). For the analysis, all areas of light scattering on each image, showing the presence of biofilm, were selected, and the software analyzed the values in square millimeters.
For the microbiologic analysis, after 72 hours, each plastic tooth was washed by immersion in phosphate-buffered saline solution to remove the brain-heart infusion broth and placed in a microcentrifuge tube with 1 mL of phosphate-buffered saline solution, that was subsequently sealed and vortexed for 1 minute and, after that, challenged with a low-output ultrasonication to unbind the biofilm. One hundred-microliter aliquots were added to the wells of a 96-well plate for serial dilution and streaking on square brain-heart infusion agar plates. The plates were incubated by using standard aerobic conditions at 37°C for 24 hours so that the colony-forming units could be counted according to the method of Jett et al. After we counted the colony-forming units, we computed the mean values from samples in each group.
For comparison and validation of the image area analysis, the data from both experiments were compared.
Statistical analysis
Medians and means for bacterial counts and biofilm areas with the corresponding standard deviations were calculated. The mean bacterial counts (in colony-forming units per milliliter) and the biofilm area determination (in square millimeters) from teeth with steel wire ligation, self-ligating brackets, and elastomeric rings were tested for significant differences by using analysis of variance (ANOVA) followed by Tukey tests. P <0.05 was considered statistically significant. Statistical comparisons between means were performed with software (Origin, version 8.5; OriginLab, Northampton, Mass).
Results
Biofilm was detected on all orthodontic brackets by fluorescence and OCT images. The addition of 1 mL of a suspension containing 10 9 cells of S mutans into the petri dish with brain-heart infusion broth and the samples followed by 3 days of incubation at 37°C in aerobic conditions reliably and reproducibly produced fluorescent biofilms ( Fig 1 , A ). Irrigation with 10 mL of saline solution did not affect significantly the fluorescence signal when analyzed by fluorescence microscopy. Figure 1 , A , shows the fluorescent image that confirms the biofilm presence on the brackets, and Figure 1 , B , shows a bracket with an elastomeric ring after the biofilm challenge.
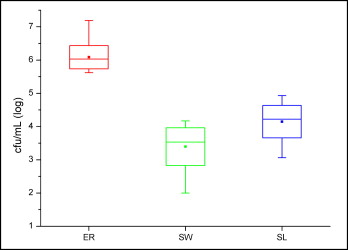
The microbiologic analysis showed a significant difference on biofilm formation depending on the ligating method. Brackets ligated with elastomeric rings collected more S mutans biofilm than did self-ligating brackets and steel wire ligation. Figure 2 gives the data obtained from the biofilms removed from each type of ligation. The initial bacterial burden did vary between individual brackets. For elastomeric rings, the mean value was 6.09 log colony-forming units per milliliter (range, 7.19-5.62). Self-ligating brackets had less biofilm compared with elastomeric rings, an average of 4.14 log colony-forming units per milliliter (range, 4.93-3.09). However, steel wire brackets showed still less biofilm, with a mean value of colony-forming units per milliliter of 3.4 log (range, 4.17-2).