Introduction
Vitamin D is responsible for the regulation of certain genes at the transcription level, via interaction with the vitamin D receptor, and influences host immune responses and aspects of bone development, growth, and homeostasis. Our aim was to investigate the association of T a q I vitamin D receptor gene polymorphism with external apical root resorption during orthodontic treatment.
Methods
Our subjects were 377 patients with Class II Division 1 malocclusion, divided into 3 groups: (1) 160 with external apical root resorption ≤1.43 mm, (2) 179 with external apical root resorption >1.43 mm), and (3) 38 untreated subjects. External apical root resorption of the maxillary incisors was evaluated on periapical radiographs taken before and after 6 months of treatment. After DNA collection and purification, vitamin D receptor T a q I polymorphism analysis was performed by polymerase chain reaction-restriction fragment length polymorphism. Univariate and multivariate analyses were performed to verify the association of clinical and genetic variables with external apical root resorption ( P <0.05).
Results
There was a higher proportion of external apical root resorption in orthodontically treated patients compared with the untreated subjects. In patients orthodontically treated, age higher than 14 years old, initial size of the maxillary incisor root superior to 30 mm, and premolar extraction were associated with increased external apical root resorption. Genotypes containing the C allele were weakly associated with protection against external apical root resorption (CC + CT × TT [odds ratio, 0.29; 95% confidence interval, 0.07-1.23; P = 0.091]) when treated orthodontic patients were compared to untreated individuals.
Conclusions
Clinical factors and vitamin D receptor T a q I polymorphism were associated with external apical root resorption in orthodontic patients.
External apical root resorption (EARR) is a common consequence of orthodontic tooth movement. Many studies have aimed to elucidate the causal relationship between orthodontic tooth movement and root resorption, but to date this issue is poorly understood, and it is not possible to predict who will develop EARR. The clinical manifestation of EARR in orthodontic patients is highly variable. Most orthodontic patients have moderate resorption that does not harm the dentition; in others, it is severe, with an unfavorable prognosis.
Root resorption during orthodontic movement is considered to be the result of an inflammatory reaction; therefore, some researchers have called this process orthodontically induced inflammatory root resorption . Because the inflammatory extent of root resorption is mainly manifested across the root contour, it can be more specifically defined as EARR.
It is believed that EARR is not caused only by orthodontic movement, but, in patients under treatment, it can result from many variables, such as host (genetic aspects) and environmental (mechanical forces, trauma) factors. A difficulty of evaluating the causes of EARR is to separate the contributions made by genetics from environmental factors such as treatment.
It was reported that genetic factors account for at least 50% of the variation in EARR. It has been suggested that there is a strong genetic component for EARR, estimated in 70% of patients, especially when dizygotic are compared to monozygotic twins. In spite of efforts to identify the genetic component for EARR, how many and which genes contribute to the phenotype of EARR are yet largely unknown.
Vitamin D is responsible for both positive and negative control of certain genes at the level of transcription, via interaction with the vitamin D receptor and is important for skeletal growth and bone homeostasis. The human vitamin D receptor is a product of 1 gene, which is located on chromosome 12q13-14. The gene comprises 9 exons that, together with intervening introns, span approximately 63 kilobases (kb). Genome-wide analyses have identified over 100 polymorphisms in the vitamin D receptor gene. Polymorphisms refer to the existence of 2 or more alleles at a given locus; if such alleles occur at a frequency of more than 1% in a population, the locus is said to be polymorphic. Single nucleotide polymorphisms are the most common form of DNA variation in the human genome. Patterns of linkage disequilibrium in the vitamin D receptor gene were proposed for a Canadian population, and 2 linkage disequilibrium blocks are believed to represent the whole gene. Block 1 is located toward the 5′ end and spans roughly 8.4 kb, and block 2 is located toward the 3′ end of the vitamin D receptor and spans approximately 5.8 kb. Near the 3′ untranslated region, there is a single nucleotide polymorphism identified by a restriction site for T a q I enzyme in vitamin D receptor exon 9 (rs731236). This single nucleotide polymorphism might represent the second linkage disequilibrium block. Alleles of this polymorphism are in linkage disequilibrium with other polymorphisms in the same block and are linked and inherited together. Allelic variations in this region could be responsible for messenger RNA stability and differences in translational efficiency, resulting in changes in cellular expression of vitamin D receptor. This polymorphism has been associated with bone mass, turnover, and mineral loss and diseases such as osteoarthritis and periodontal disease. However, no study has investigated vitamin D receptor gene polymorphisms and their association with EARR.
The aim of this study was to investigate the association of T a q I vitamin D receptor enzyme polymorphism (rs731236, exon 9) and clinical variables with EARR in orthodontic patients.
Material and methods
A sample of 377 unrelated white patients of both sexes (mean age, 14.9 years; range, 8-21 years) was selected between 2005 and 2009. Of these, 339 patients had a Class II Division 1 malocclusion and were orthodontically treated with edgewise or straight-wire techniques, and 38 patients had a Class II Division 1 malocclusion but were untreated. The reason for Class II Division 1 subjects was because this type of malocclusion is the most frequent and requires more treatment ; in addition, it can lead to higher levels of EARR. The patients were selected from the Dental Clinics of the Graduate Course in Orthodontics (Bauru, São Paulo, Brazil), Dental Clinics of the Graduate Course in Orthodontics of Thum Institute of Research (Joinville, Santa Catarina, Brazil), and 2 private orthodontic clinics (Curitiba, Paraná, Brazil) ( Table I ). Although all subjects were white, the Brazilian white population is heterogeneous. Recent articles have not recommended grouping Brazilians into ethnic groups based on color, race, and geographic origin because Brazilians classified as white or black have significantly overlapping genotypes, probably due to miscegenation.
| Group 1 n = 160 (%) |
Group 2 n = 179 (%) |
Group 3 n = 38 (%) |
P value | |
|---|---|---|---|---|
| Sex (n) ∗ | ||||
| Female | 86 (53.7%) | 99 (55.3%) | 23 (60.5%) | 0.695 |
| Male | 74 (46.3%) | 80 (44.7%) | 15 (39.5%) | |
| Age (y) † | ||||
| Mean ± SD (range) | 14.50 ± 3.01 (8-21) | 15.33 ± 2.64 (9.9-20) | 16.46 ± 1.93 (11-19) | <0.001 |
The subjects completed personal, medical, and dental history questionnaires and, with the protocol approved by the institutional review board of Pontifical Catholic University of Paraná, signed a consent form after being advised of the nature of the study (approved by the ethical committee in research at Pontificia Unieversidade Católica do Paraná, protocol number 546/05). The patients could not have chronic usage of anti-inflammatory drugs, human immunodeficiency virus infection, and immunosuppressive chemotherapy history of any disease known to severely compromise immune function, current pregnancy or lactation, oral trauma, parafunctional behavior, endodontic treatment, or extensive caries lesions.
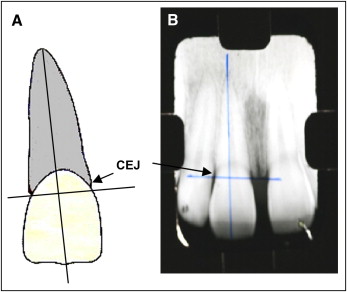
The periapical x-rays of the maxillary central incisors with the longer roots (reference teeth) were taken before treatment and 6 months after the beginning of treatment. The evaluation method included measuring the root and crown lengths directly from the radiographs. The root apex, incisal edge, and cementoenamel junction of each tooth were marked on the x-rays on a light table. The longitudinal axis of each tooth was constructed from the root apex to the incisal edge following the root canal as accurately as possible. A perpendicular axis was then projected to the longitudinal axis from the mesial to the distal cementoenamel junction sides. The crown length was measured from the incisal edge to the projected cementoenamel junction, and the root length from the projected cementoenamel junction to the root apex ( Fig 1 ). The differences between the 2 measurements indicate the EARR. A correction factor was calculated by using the following formula: correction factor = C1/C2 ( C1 , crown length before treatment; C2 , crown length 6 months after starting treatment). Then EARR was calculated with the following formula: EARR = R1 – (R2 × CF) ( R1 , root length before treatment; R2 , root length 6 months after treatment start; CF , correction factor). EARR was also expressed as a percentage of the original root length: EARR × 100/R1. Only teeth with complete root formation were considered for investigation. Any distortions between the pretreatment and follow-up radiographic images were corrected by using the crown length registration, assuming crown length to be unchanging over the observation period. The EARR was evaluated by 1 examiner (M.L.S.S.N.F). The radiographs were examined over a light table, and the measurements were made with a fine-tip digital caliper with accuracy up to 0.02 mm (electronic digital vernier caliper; Utustools Professional, Santiago, Chile).
A receiver operating characteristic curve was constructed to verify the cutoff point based on the sample data distribution for EARR, and the value of 1.43 mm was obtained. Then, the sample was divided into 3 groups: (1) 160 treated patients with EARR 1.43 mm or less, (2) 179 treated patients with EARR greater than 1.43 mm, and (3) 38 untreated subjects.
The following parameters were evaluated in the treated patients: age, sex, initial size of the root of the reference tooth, premolar extraction, use of pendulum appliance, rapid palatal expansion, and use of elastics ( Table II ).
| Variable | n | EARR | P value ∗ (univariate) n = 339 |
P value † (multivariate) n = 339 |
Odds ratio | 95% Confidence interval | |
|---|---|---|---|---|---|---|---|
| ≤1.43 mm n = 160 (47.2%) |
>1.43 mm n = 179 (52.8%) |
||||||
| Age (y) ‡ | |||||||
| ≤14 | 147 | 80 (54.4) | 67 (45,6) | ||||
| >14 | 192 | 80 (41.7) | 112 (58.3) | 0.021 | 0.022 | 1.69 | 1.08-2.66 |
| Sex | |||||||
| Female | 185 | 86 (46.5) | 99 (53.5) | ||||
| Male | 154 | 74 (48.1) | 80 (51.9) | 0.827 | |||
| Initial root (mm) § | |||||||
| <30 | 258 | 133 (51.6) | 125 (48.4) | ||||
| ≥30 | 81 | 27 (33.3) | 54 (66.7) | 0.005 | 0.002 | 2.34 | 1.36-4.03 |
| Premolar extraction | |||||||
| No | 291 | 143 (49.1) | 148 (50.9) | ||||
| Yes | 48 | 17 (35.4) | 31 (64.6) | 0.087 | 0.052 | 1.94 | 0.99-3.78 |
| Elastics | |||||||
| No | 276 | 133 (48.2) | 143 (51.8) | ||||
| Yes | 63 | 42 (42.9) | 36 (57.1) | 0.486 | |||
| Rapid palatal expansion | |||||||
| No | 285 | 130 (45.6) | 155 (54.4) | ||||
| Yes | 54 | 30 (55.6) | 24 (44.4) | 0.185 | |||
| Pendulum | |||||||
| No | 252 | 113 (44.8) | 139 (55.2) | ||||
| Yes | 87 | 47 (54) | 40 (46) | 0.171 | |||
| Genotype | |||||||
| TT | 58 | 33 (56.9) | 25 (43.1) | ||||
| TC + CC | 274 | 124 (45.3) | 150 (54.7) | 0.114 | 0.104 | 1.64 | 0.90-2.97 |
† regression logistic model, Wald test, P <0.05 (variables were included when P <0.20 in univariate analysis).
‡ cutoff point (14 years) suggested by receiver operating characteristic curve (0.574, P = 0.017).
§ cutoff point (30 mm) suggested by receiver operating characteristic curve (0.620, P <0.001).
Cells were obtained by a mouthwash with 3% glucose solution for 1 minute and by scraping the oral mucosa with a sterile spatula. DNA was extracted from epithelial oral cells with ammonium acetate (10 mol/L) and EDTA (1 mmol/L).
The following primer pair was used for polymerase chain reaction amplification of genomic DNA samples: (F – 5′CAG AGC ATG GAC AGG GAG CAA G 3′ and R – 5′GGA TGT ACG TCT GCA GTG TG 3′). Reaction conditions and cycling parameters were as follows: 1 μL of the genomic DNA was used for polymerase chain reaction amplification in a reaction mixture containing 22.5 μL GoTaq Green Master Mix (Promega, Madison, Wis) and 0.7 μL of each 25 μmol/L primer. The reactions were performed in a thermal cycler (T-512; Techne, Chelmsford, UK) and consisted of an initial denaturation step at 95°C for 5 minutes, followed by 37 cycles at 95°C for 1 minute, 55°C for 1 minute, 72°C for 1 minute, and a final extension at 72°C for 7 minutes. The restriction fragment length polymorphism technique was performed in a final reaction volume of 20 μL, by using 1 unit of T a q I (T↓CGA) (Invitrogen Life Technologies, Carlsbad, Calif) and a 10-μL aliquot of polymerase chain reaction products, digested at 65°C overnight. The digested products were separated in 10% polyacrylamide gel electrophoresis stained by silver. The genotypes were determined by comparing the restriction fragment length polymorphism band patterns with a 1-kb-plus DNA ladder (Invitrogen Life Technologies). The restriction fragment length polymorphism is formed by a single base transition (T a q I enzyme) at codon 352 in exon 9 of the vitamin D receptor gene that creates a T a q I restriction site. The alleles that result from the cleavage of T a q I are designated “C” (T a q I site present, with 2 fragments: 293 base pairs and 47 base pairs) or “T” (T a q I site absent, with a fragment: 340 base pairs).
Statistical analysis
The results observed in the study were expressed in means and standard deviations (quantitative variable) or frequencies and percentages (qualitative variable). To evaluate the association between 2 qualitative variables, the chi-square test or the Fisher exact test was used. Comparisons between the groups in relation to the quantitative variables were made with analysis of variance (ANOVA) with 1 factor and the least significant difference test for multiple comparisons. Adjustments of the receiver operating characteristic curve were made for EARR, age, and initial root length to determine cutoff points associated with EARR. The unpaired t test was used to compare EARR, age, and initial root length between the groups. For the multivariate analysis, the logistic regression model and the Wald test were used. Values of P <0.05 indicated statistical significance. Data were organized in an Excel spread sheet (Microsoft, Redmond, Wash) and analyzed with the computational program Statistica (version 8.0; StatSoft, Tulsa, Okla).
Results
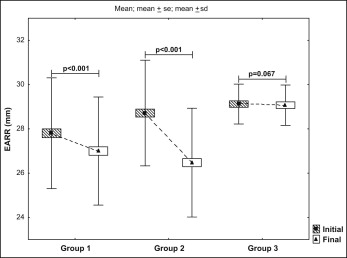
We observed a higher proportion of EARR in the treated patients (EARR ≤1.43 mm, 0.81 mm; EARR >1.43 mm, 2.24 mm) compared with the untreated subjects (EARR, 0.05 mm). The clinical impact of orthodontic appliances on root resorption is shown in Figure 2 .
Regarding the treated patients, no statistically significant differences were observed between the groups in relation to sex, pendulum appliance, rapid palatal expansion, or elastics. A statistically significant difference was found between the groups regarding age ( P = 0.021) and initial size of the root of the reference tooth ( P = 0.005) in the univariate analysis. After the multivariate analysis, age ( P = 0.022), initial size of the root of the reference tooth ( P = 0.002), and premolar extractions ( P = 0.052) were associated with EARR ( Table II ).
Considering the study’s single nucleotide polymorphism, the genotype distribution was not consistent with the assumption of Hardy-Weinberg equilibrium either in the control group or for the whole sample. Differences were observed in vitamin D receptor T a q I polymorphism genotype frequency ( P = 0.051), but not in allele distribution ( P = 0.455) between the groups ( Table III ). However, when the treated patients were analyzed vs the untreated subjects, we observed a weak protection effect of allele C against EARR (CC + CT × TT [odds ratio, 0.29; 95% confidence interval, 0.07-1.23; P = 0.091]).



